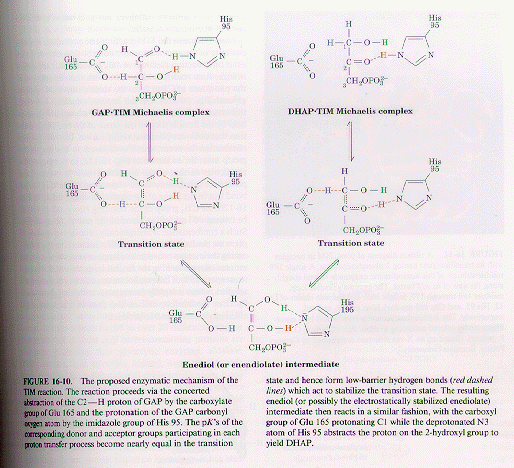
Figure 1. From Voet & Voet
Triose phosphate isomerase (TIM) catalyses the interconversion of dihydroxyacetone phosphate (DHAP) and D-glyceraldehyde-3-phosphate (GAP) in the fifth reaction of glycolysis. It is a reaction similar to that catalyzed by phosphoglucose isomerase in the second step of glycolysis (see figure 1). Glycolysis is the metabolic pathway by which glucose is converted to pyruvate. Pyruvate then continues under anaerobic conditions to homolactic or alcoholic fermentation and under aerobic conditions to the TCA cycle. Glycolysis can be considered to occur in two stages: Stage I The preparatory stage in which the hexose glucose is phosphorylated and cleaved to yield two molecules of the triose GAP, using two molecules of ATP. Stage II converts the two molecules of GAP to pyruvate and generates four ATPs. The reaction catalyzed by TIM is the final reaction of stage I and a crucial isomerization in the path of glycolysis because only GAP can continue to stage II.

Figure 1. From Voet & Voet
TIM catalyzes the reaction via concerted general acid-base catalysis involving low barrier hydrogen bonds, to bind the transition state complex more tightly than the substrate. The low-barrier hydrogen bonds occur between Glu167 and C2÷H and between His95 and the carbonyl atom in the transition state. The positively charged Lys13 is thought to electrostatically stabilize the negatively charged transition state (see figure 2). When substrate binds to TIM a conserved 10-residue loop closes over the active site like a hinged lid (see figure 3). A four-residue segment of this loop makes a hydrogen bond with the phosphate group of the substrate. Mutagenic excision of these four residues does not significantly distort the protein so that substrate binding is not greatly impaired. The catalytic power of the mutant enzyme is, nevertheless, reduced by a factor of 100,000. This indicates that the loop closure preferentially stabilizes the enzymatic reaction's enediol-like transition state. TIM was the first protein known to contain a a / b barrel, a folding motif in which eight b strands alternating in sequence with eight a helices (see figure 4). The active sites of all known a / b barrel enzymes are located in the mouth of the barrel at the end that contains the C-terminal ends of the bstrands, although there is no obvious structural rationale for this (Voet & Voet, 1995).
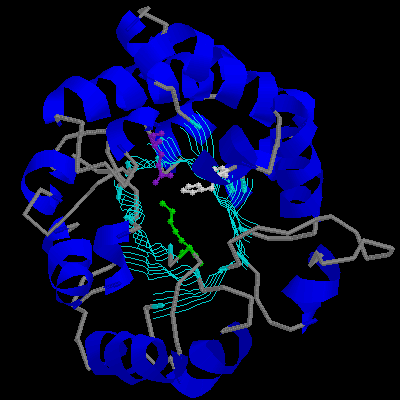

Figure 2. The important active-site residues shown down the axis of the a / b barrel. Lys13, His95, and Glu167, green, white, and purple (right). A close up of the same residues (left).
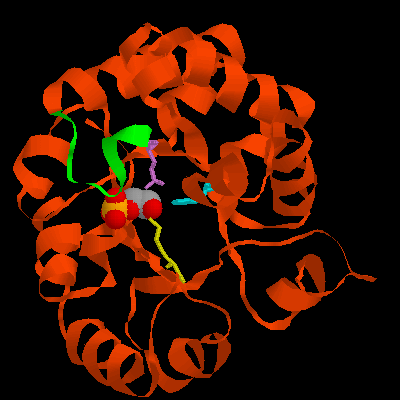
Figure 3. A ribbon diagram of L. mexicana TIM. The enzyme's flexible loop, residues 167-179, is green and the side chains of Lys13, His95, and Glu167 are yellow, cyan, and violet, respectively. The ligand is shoen according to atom type (C, grey; O, red; P, yellow).
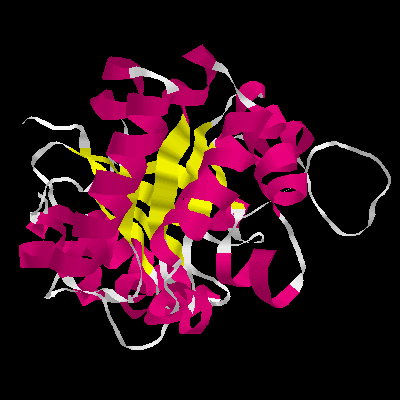
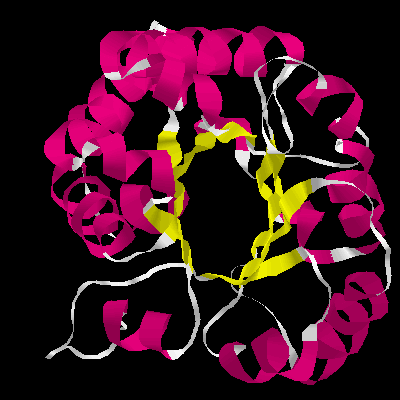
Figure 4. The a / b barrel structure of TIM side view (right) top view (left). Each subunit is composed of alternating b- strands and a- helices. The eight parallel b- strands form an inner cylinder. This b- barrel is shielded from the solvent by eight helices, running antiparallel to the strands of the inner barrel.